Expo
view channel
view channel
view channel
view channel
view channel
view channel
view channel
RadiographyMRIUltrasoundNuclear Medicine
Imaging ITIndustry News
Events

- AI Detection Tool Improves Identification of Lobular Breast Cancer
- New Contrast Agent Enables Low-Dose X-Ray Joint Imaging
- AI Boosts Breast Cancer Detection and Cuts Screening Workload
- AI Tool Predicts Breast Cancer Risk Years Ahead Using Routine Mammograms
- Routine Mammograms Could Predict Future Cardiovascular Disease in Women
- AI Body Composition MRI Analysis Predicts Cardiometabolic Disease Risk
- Blood-Brain Barrier Imaging Adds Risk Insight to Standard Stroke MRI
- AI MRI Tool Quantifies Muscle Fat to Assess Cardiometabolic Risk
- Advanced MRI Visualizes CSF Motion Changes After Mild Traumatic Brain Injury
- MRI Tool Enables Long-Term Tracking of Transplanted Cardiac Cells
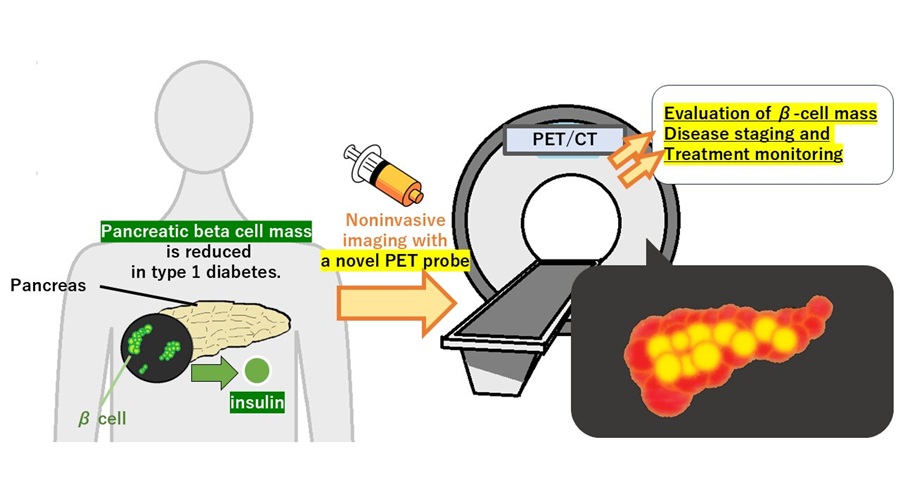
- PET Tracer Enables Noninvasive Measurement of Beta Cell Mass
- New Imaging Tool Sheds Light on Tumor Fat Metabolism
- Radiopharmaceutical Molecule Marker to Improve Choice of Bladder Cancer Therapies
- Cancer “Flashlight” Shows Who Can Benefit from Targeted Treatments
- PET Imaging of Inflammation Predicts Recovery and Guides Therapy After Heart Attack
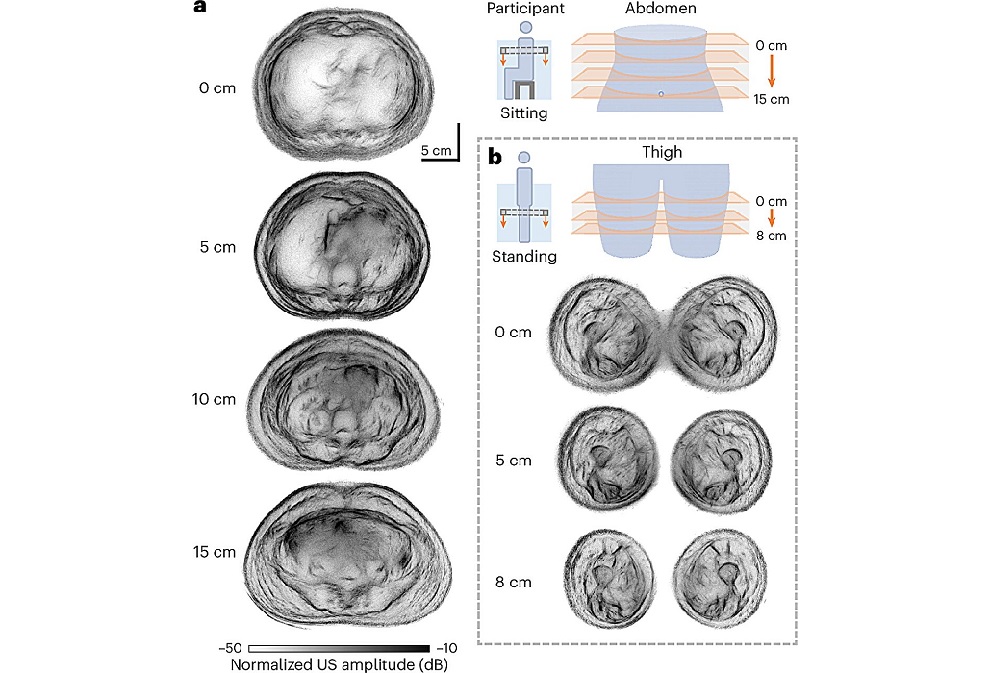
- Whole Cross-Section Ultrasound System Enables Operator-Independent Imaging
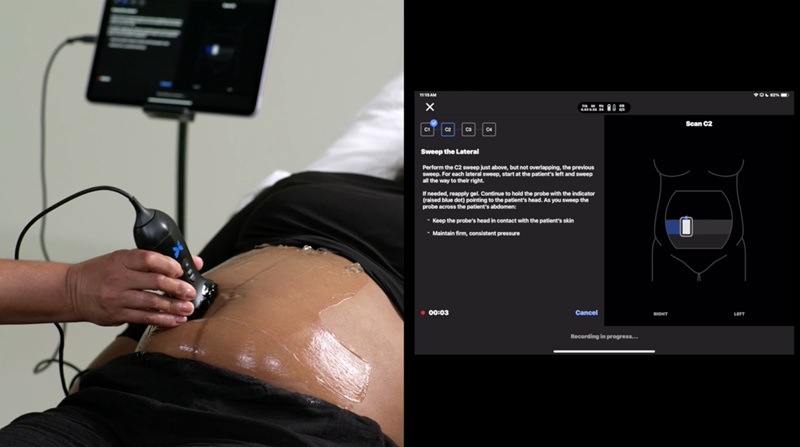
- New Ultrasound AI Tool Supports Rapid Prenatal Assessment
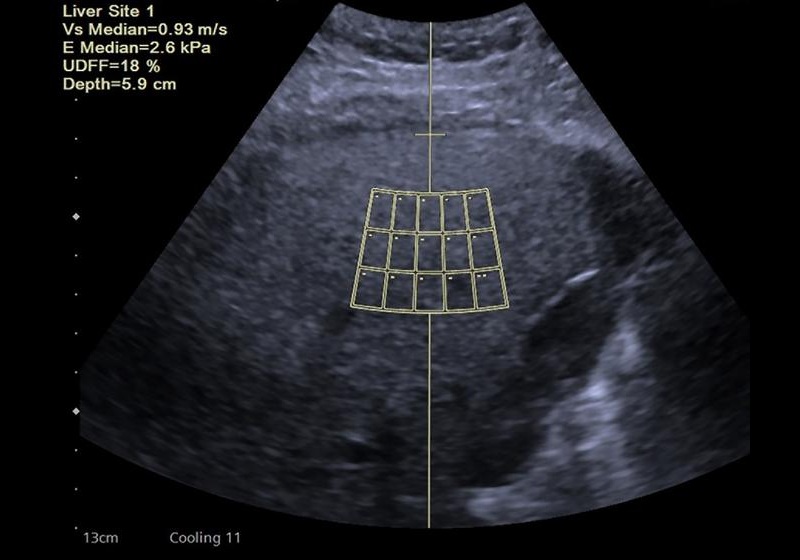
- New Consensus Standardizes Ultrasound-Based Fatty Liver Assessment
- Groundbreaking Technology to Enhance Precision in Emergency and Critical Care
- Reusable Gel Pad Made from Tamarind Seed Could Transform Ultrasound Examinations
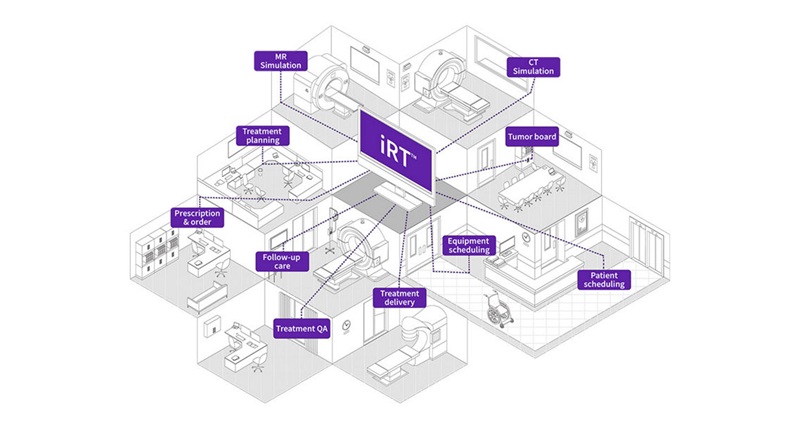
- New Proton Therapy Platform Integrates into Existing Radiotherapy Departments
- 3D-Printed Intraoral Device Enhances Head and Neck Radiotherapy Accuracy
- Molecular Imaging Agent Shows Promise for Endometriosis Detection and Monitoring
- Automated AI Tool Detects Early Pancreatic Cancer on Routine CT
- Routine Cardiac CT Enhanced to Predict Heart Failure Risk
- Breast Imaging Software Enhances Visualization and Tissue Characterization in Challenging Cases
- New Google Cloud Medical Imaging Suite Makes Imaging Healthcare Data More Accessible
- Global AI in Medical Diagnostics Market to Be Driven by Demand for Image Recognition in Radiology
- AI-Based Mammography Triage Software Helps Dramatically Improve Interpretation Process
- Artificial Intelligence (AI) Program Accurately Predicts Lung Cancer Risk from CT Images
- Nuclear Medicine Set for Continued Growth Driven by Demand for Precision Diagnostics
- GE HealthCare and NVIDIA Collaboration to Reimagine Diagnostic Imaging
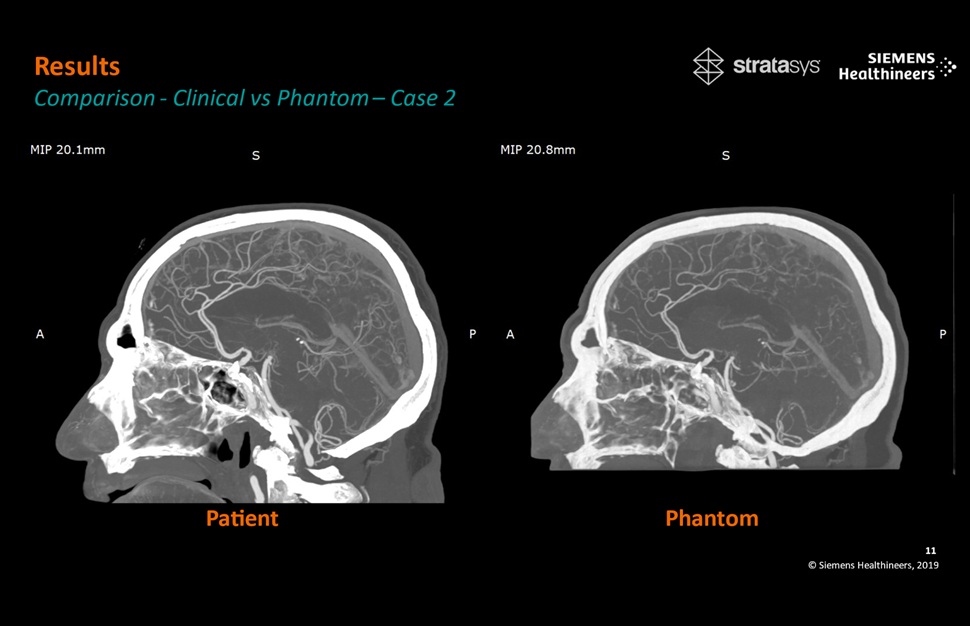
- Patient-Specific 3D-Printed Phantoms Transform CT Imaging
- Siemens and Sectra Collaborate on Enhancing Radiology Workflows
- Bracco Diagnostics and ColoWatch Partner to Expand Availability CRC Screening Tests Using Virtual Colonoscopy

 Expo
Expo
- AI Detection Tool Improves Identification of Lobular Breast Cancer
- New Contrast Agent Enables Low-Dose X-Ray Joint Imaging
- AI Boosts Breast Cancer Detection and Cuts Screening Workload
- AI Tool Predicts Breast Cancer Risk Years Ahead Using Routine Mammograms
- Routine Mammograms Could Predict Future Cardiovascular Disease in Women
- AI Body Composition MRI Analysis Predicts Cardiometabolic Disease Risk
- Blood-Brain Barrier Imaging Adds Risk Insight to Standard Stroke MRI
- AI MRI Tool Quantifies Muscle Fat to Assess Cardiometabolic Risk
- Advanced MRI Visualizes CSF Motion Changes After Mild Traumatic Brain Injury
- MRI Tool Enables Long-Term Tracking of Transplanted Cardiac Cells
- PET Tracer Enables Noninvasive Measurement of Beta Cell Mass
- New Imaging Tool Sheds Light on Tumor Fat Metabolism
- Radiopharmaceutical Molecule Marker to Improve Choice of Bladder Cancer Therapies
- Cancer “Flashlight” Shows Who Can Benefit from Targeted Treatments
- PET Imaging of Inflammation Predicts Recovery and Guides Therapy After Heart Attack
- Whole Cross-Section Ultrasound System Enables Operator-Independent Imaging
- New Ultrasound AI Tool Supports Rapid Prenatal Assessment
- New Consensus Standardizes Ultrasound-Based Fatty Liver Assessment
- Groundbreaking Technology to Enhance Precision in Emergency and Critical Care
- Reusable Gel Pad Made from Tamarind Seed Could Transform Ultrasound Examinations
- New Proton Therapy Platform Integrates into Existing Radiotherapy Departments
- 3D-Printed Intraoral Device Enhances Head and Neck Radiotherapy Accuracy
- Molecular Imaging Agent Shows Promise for Endometriosis Detection and Monitoring
- Automated AI Tool Detects Early Pancreatic Cancer on Routine CT
- Routine Cardiac CT Enhanced to Predict Heart Failure Risk
- Breast Imaging Software Enhances Visualization and Tissue Characterization in Challenging Cases
- New Google Cloud Medical Imaging Suite Makes Imaging Healthcare Data More Accessible
- Global AI in Medical Diagnostics Market to Be Driven by Demand for Image Recognition in Radiology
- AI-Based Mammography Triage Software Helps Dramatically Improve Interpretation Process
- Artificial Intelligence (AI) Program Accurately Predicts Lung Cancer Risk from CT Images
- Nuclear Medicine Set for Continued Growth Driven by Demand for Precision Diagnostics
- GE HealthCare and NVIDIA Collaboration to Reimagine Diagnostic Imaging
- Patient-Specific 3D-Printed Phantoms Transform CT Imaging
- Siemens and Sectra Collaborate on Enhancing Radiology Workflows
- Bracco Diagnostics and ColoWatch Partner to Expand Availability CRC Screening Tests Using Virtual Colonoscopy